Iron »
PDB 2d3y-2e1q »
2d40 »
Iron in PDB 2d40: Crystal Structure of Z3393 From Escherichia Coli O157:H7
Enzymatic activity of Crystal Structure of Z3393 From Escherichia Coli O157:H7
All present enzymatic activity of Crystal Structure of Z3393 From Escherichia Coli O157:H7:
1.13.11.4;
1.13.11.4;
Protein crystallography data
The structure of Crystal Structure of Z3393 From Escherichia Coli O157:H7, PDB code: 2d40
was solved by
M.A.Adams,
Z.Jia,
Montreal-Kingston Bacterial Structural Genomicsinitiative (Bsgi),
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 34.50 / 2.41 |
| Space group | P 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 54.134, 76.075, 85.464, 114.08, 94.93, 108.11 |
| R / Rfree (%) | 17.3 / 24.9 |
Iron Binding Sites:
The binding sites of Iron atom in the Crystal Structure of Z3393 From Escherichia Coli O157:H7
(pdb code 2d40). This binding sites where shown within
5.0 Angstroms radius around Iron atom.
In total 4 binding sites of Iron where determined in the Crystal Structure of Z3393 From Escherichia Coli O157:H7, PDB code: 2d40:
Jump to Iron binding site number: 1; 2; 3; 4;
In total 4 binding sites of Iron where determined in the Crystal Structure of Z3393 From Escherichia Coli O157:H7, PDB code: 2d40:
Jump to Iron binding site number: 1; 2; 3; 4;
Iron binding site 1 out of 4 in 2d40
Go back to
Iron binding site 1 out
of 4 in the Crystal Structure of Z3393 From Escherichia Coli O157:H7
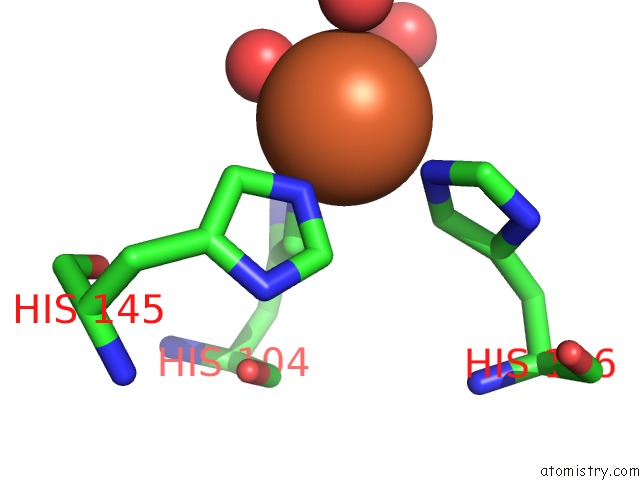
Mono view
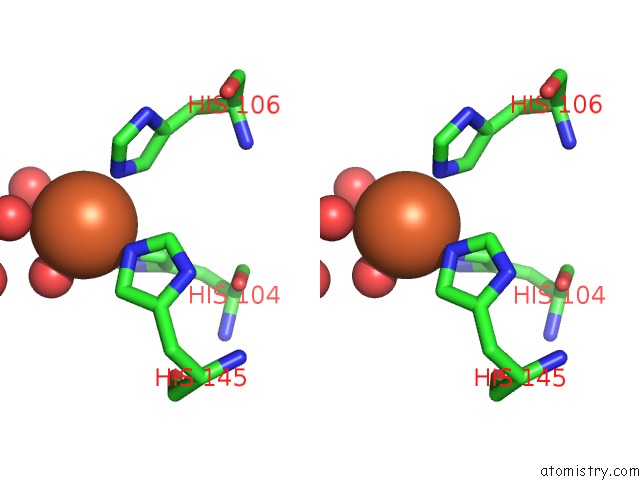
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 1 of Crystal Structure of Z3393 From Escherichia Coli O157:H7 within 5.0Å range:
|
Iron binding site 2 out of 4 in 2d40
Go back to
Iron binding site 2 out
of 4 in the Crystal Structure of Z3393 From Escherichia Coli O157:H7
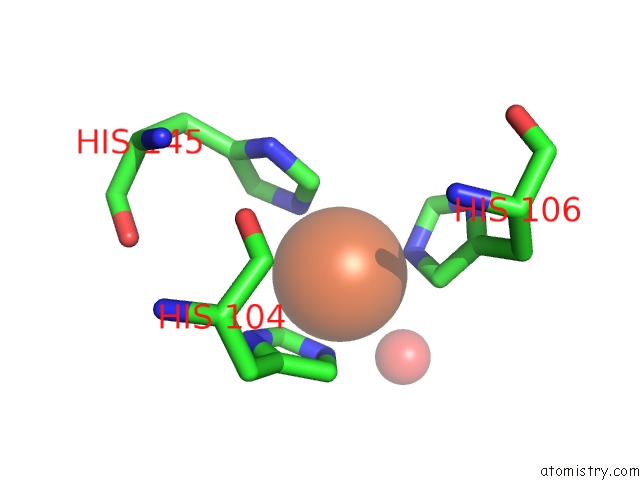
Mono view
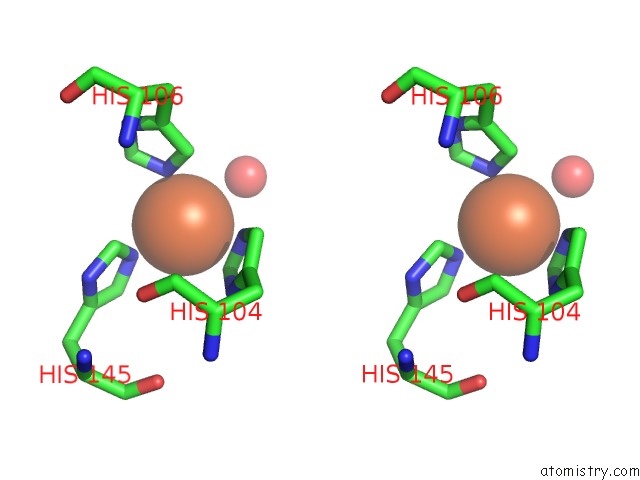
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 2 of Crystal Structure of Z3393 From Escherichia Coli O157:H7 within 5.0Å range:
|
Iron binding site 3 out of 4 in 2d40
Go back to
Iron binding site 3 out
of 4 in the Crystal Structure of Z3393 From Escherichia Coli O157:H7
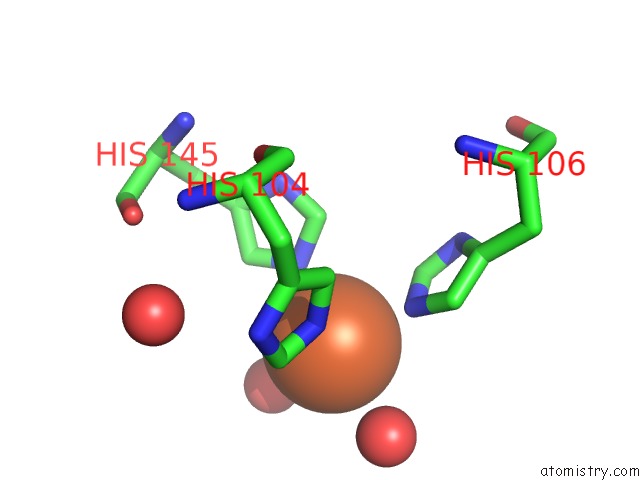
Mono view
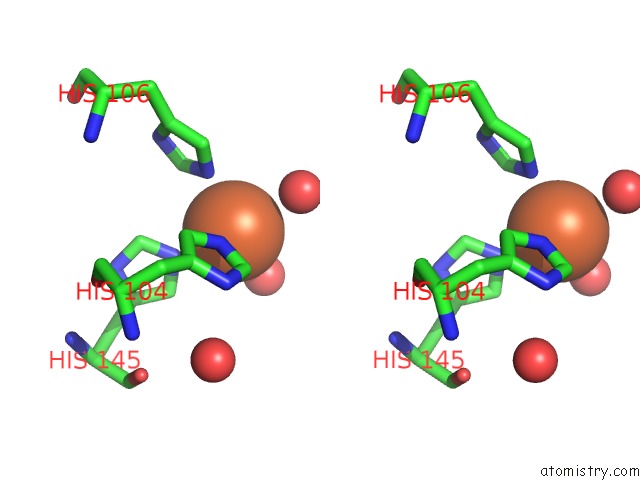
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 3 of Crystal Structure of Z3393 From Escherichia Coli O157:H7 within 5.0Å range:
|
Iron binding site 4 out of 4 in 2d40
Go back to
Iron binding site 4 out
of 4 in the Crystal Structure of Z3393 From Escherichia Coli O157:H7
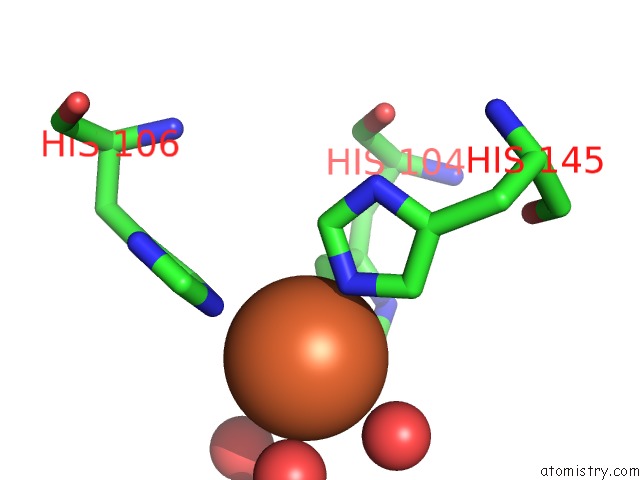
Mono view
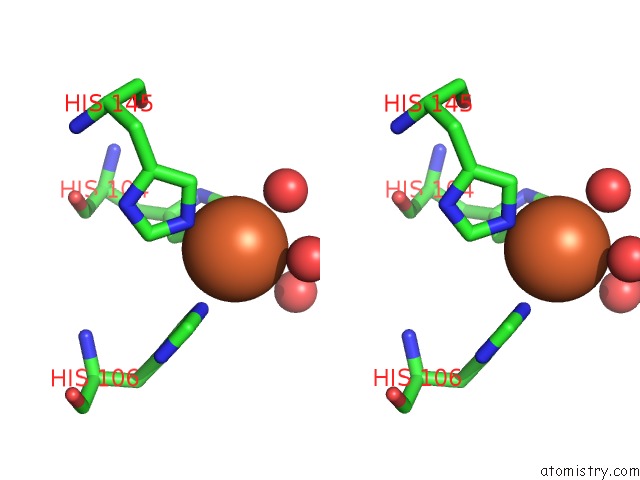
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 4 of Crystal Structure of Z3393 From Escherichia Coli O157:H7 within 5.0Å range:
|
Reference:
M.A.Adams,
V.K.Singh,
B.O.Keller,
Z.Jia.
Structural and Biochemical Characterization of Gentisate 1,2-Dioxygenase From Escherichia Coli O157:H7 Mol.Microbiol. V. 61 1469 2006.
ISSN: ISSN 0950-382X
PubMed: 16930152
DOI: 10.1111/J.1365-2958.2006.05334.X
Page generated: Sat Aug 3 20:43:21 2024
ISSN: ISSN 0950-382X
PubMed: 16930152
DOI: 10.1111/J.1365-2958.2006.05334.X
Last articles
Cl in 5SDKCl in 5SDI
Cl in 5SDJ
Cl in 5SDH
Cl in 5SDD
Cl in 5SDE
Cl in 5SDF
Cl in 5SDG
Cl in 5SDC
Cl in 5SD6