Iron »
PDB 2qbl-2r1l »
2r13 »
Iron in PDB 2r13: Crystal Structure of Human Mitoneet Reveals A Novel [2FE-2S] Cluster Coordination
Protein crystallography data
The structure of Crystal Structure of Human Mitoneet Reveals A Novel [2FE-2S] Cluster Coordination, PDB code: 2r13
was solved by
X.Hou,
R.Liu,
S.Ross,
E.J.Smart,
H.Zhu,
W.Gong,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 30.00 / 1.80 |
| Space group | I 41 2 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 58.794, 58.794, 175.306, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 15.9 / 18.8 |
Other elements in 2r13:
The structure of Crystal Structure of Human Mitoneet Reveals A Novel [2FE-2S] Cluster Coordination also contains other interesting chemical elements:
| Chlorine | (Cl) | 1 atom |
Iron Binding Sites:
The binding sites of Iron atom in the Crystal Structure of Human Mitoneet Reveals A Novel [2FE-2S] Cluster Coordination
(pdb code 2r13). This binding sites where shown within
5.0 Angstroms radius around Iron atom.
In total 2 binding sites of Iron where determined in the Crystal Structure of Human Mitoneet Reveals A Novel [2FE-2S] Cluster Coordination, PDB code: 2r13:
Jump to Iron binding site number: 1; 2;
In total 2 binding sites of Iron where determined in the Crystal Structure of Human Mitoneet Reveals A Novel [2FE-2S] Cluster Coordination, PDB code: 2r13:
Jump to Iron binding site number: 1; 2;
Iron binding site 1 out of 2 in 2r13
Go back to
Iron binding site 1 out
of 2 in the Crystal Structure of Human Mitoneet Reveals A Novel [2FE-2S] Cluster Coordination
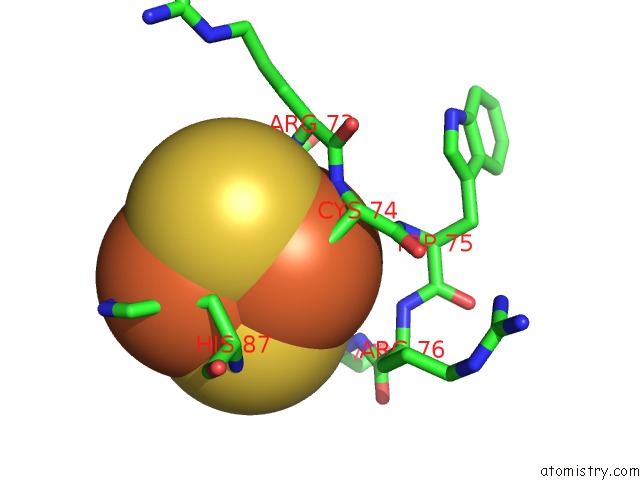
Mono view
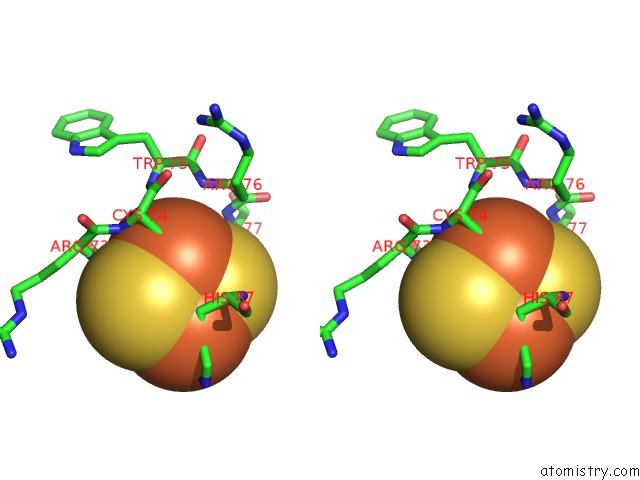
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 1 of Crystal Structure of Human Mitoneet Reveals A Novel [2FE-2S] Cluster Coordination within 5.0Å range:
|
Iron binding site 2 out of 2 in 2r13
Go back to
Iron binding site 2 out
of 2 in the Crystal Structure of Human Mitoneet Reveals A Novel [2FE-2S] Cluster Coordination
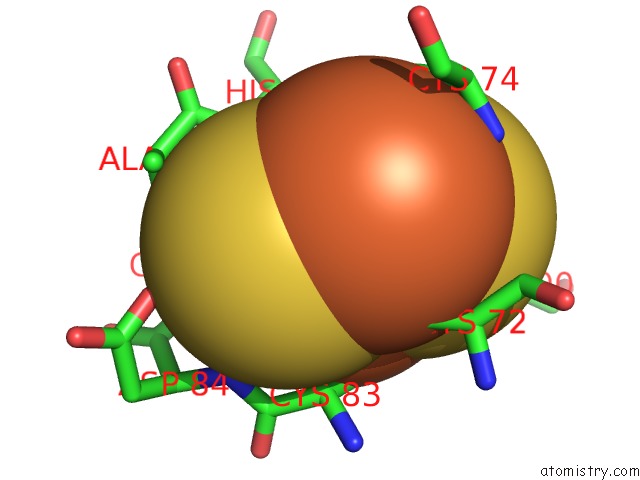
Mono view
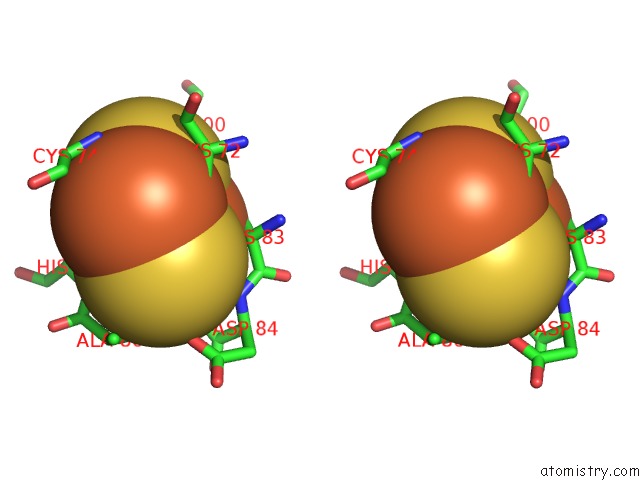
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 2 of Crystal Structure of Human Mitoneet Reveals A Novel [2FE-2S] Cluster Coordination within 5.0Å range:
|
Reference:
X.Hou,
R.Liu,
S.Ross,
E.J.Smart,
H.Zhu,
W.Gong.
Crystallographic Studies of Human Mitoneet J.Biol.Chem. V. 282 33242 2007.
ISSN: ISSN 0021-9258
PubMed: 17905743
DOI: 10.1074/JBC.C700172200
Page generated: Sun Aug 4 01:51:43 2024
ISSN: ISSN 0021-9258
PubMed: 17905743
DOI: 10.1074/JBC.C700172200
Last articles
F in 7PY4F in 7PX6
F in 7PVU
F in 7PRV
F in 7PRW
F in 7PMQ
F in 7PRX
F in 7PPH
F in 7PRM
F in 7PQV