Iron »
PDB 3jbt-3k9y »
3k0v »
Iron in PDB 3k0v: Removal of Sugars and Sugars-Like Molecules From the Solution By C- Lobe of Lactoferrin: Crystal Structure of the Complex of C-Lobe with Beta-D-Glucopyranosyl-(1->4)-Beta-D-Galactopyranosyl-(1->4)-Alpha-D- Glucopyranose at 1.9 A Resolution
Protein crystallography data
The structure of Removal of Sugars and Sugars-Like Molecules From the Solution By C- Lobe of Lactoferrin: Crystal Structure of the Complex of C-Lobe with Beta-D-Glucopyranosyl-(1->4)-Beta-D-Galactopyranosyl-(1->4)-Alpha-D- Glucopyranose at 1.9 A Resolution, PDB code: 3k0v
was solved by
R.Mir,
G.Vikram,
M.Sinha,
N.Singh,
S.Sharma,
P.Kaur,
T.P.Singh,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 62.00 / 1.91 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 61.810, 50.134, 65.545, 90.00, 107.10, 90.00 |
| R / Rfree (%) | 20.7 / 23.7 |
Other elements in 3k0v:
The structure of Removal of Sugars and Sugars-Like Molecules From the Solution By C- Lobe of Lactoferrin: Crystal Structure of the Complex of C-Lobe with Beta-D-Glucopyranosyl-(1->4)-Beta-D-Galactopyranosyl-(1->4)-Alpha-D- Glucopyranose at 1.9 A Resolution also contains other interesting chemical elements:
| Zinc | (Zn) | 2 atoms |
Iron Binding Sites:
The binding sites of Iron atom in the Removal of Sugars and Sugars-Like Molecules From the Solution By C- Lobe of Lactoferrin: Crystal Structure of the Complex of C-Lobe with Beta-D-Glucopyranosyl-(1->4)-Beta-D-Galactopyranosyl-(1->4)-Alpha-D- Glucopyranose at 1.9 A Resolution
(pdb code 3k0v). This binding sites where shown within
5.0 Angstroms radius around Iron atom.
In total only one binding site of Iron was determined in the Removal of Sugars and Sugars-Like Molecules From the Solution By C- Lobe of Lactoferrin: Crystal Structure of the Complex of C-Lobe with Beta-D-Glucopyranosyl-(1->4)-Beta-D-Galactopyranosyl-(1->4)-Alpha-D- Glucopyranose at 1.9 A Resolution, PDB code: 3k0v:
In total only one binding site of Iron was determined in the Removal of Sugars and Sugars-Like Molecules From the Solution By C- Lobe of Lactoferrin: Crystal Structure of the Complex of C-Lobe with Beta-D-Glucopyranosyl-(1->4)-Beta-D-Galactopyranosyl-(1->4)-Alpha-D- Glucopyranose at 1.9 A Resolution, PDB code: 3k0v:
Iron binding site 1 out of 1 in 3k0v
Go back to
Iron binding site 1 out
of 1 in the Removal of Sugars and Sugars-Like Molecules From the Solution By C- Lobe of Lactoferrin: Crystal Structure of the Complex of C-Lobe with Beta-D-Glucopyranosyl-(1->4)-Beta-D-Galactopyranosyl-(1->4)-Alpha-D- Glucopyranose at 1.9 A Resolution
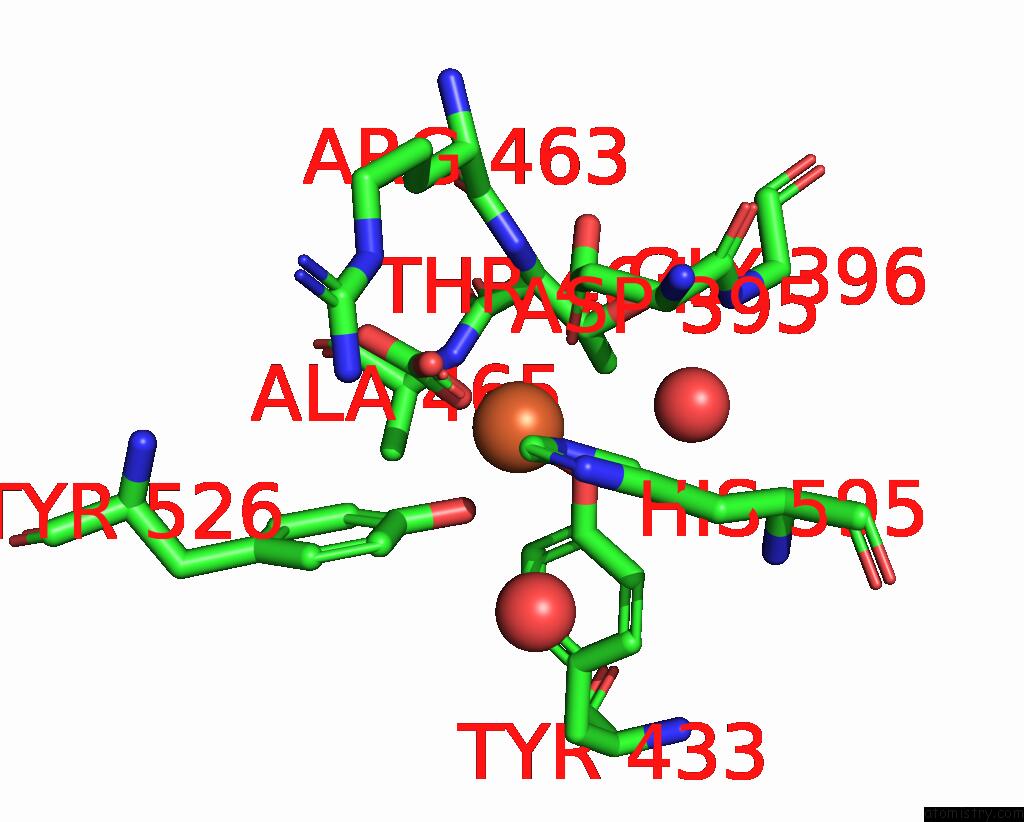
Mono view
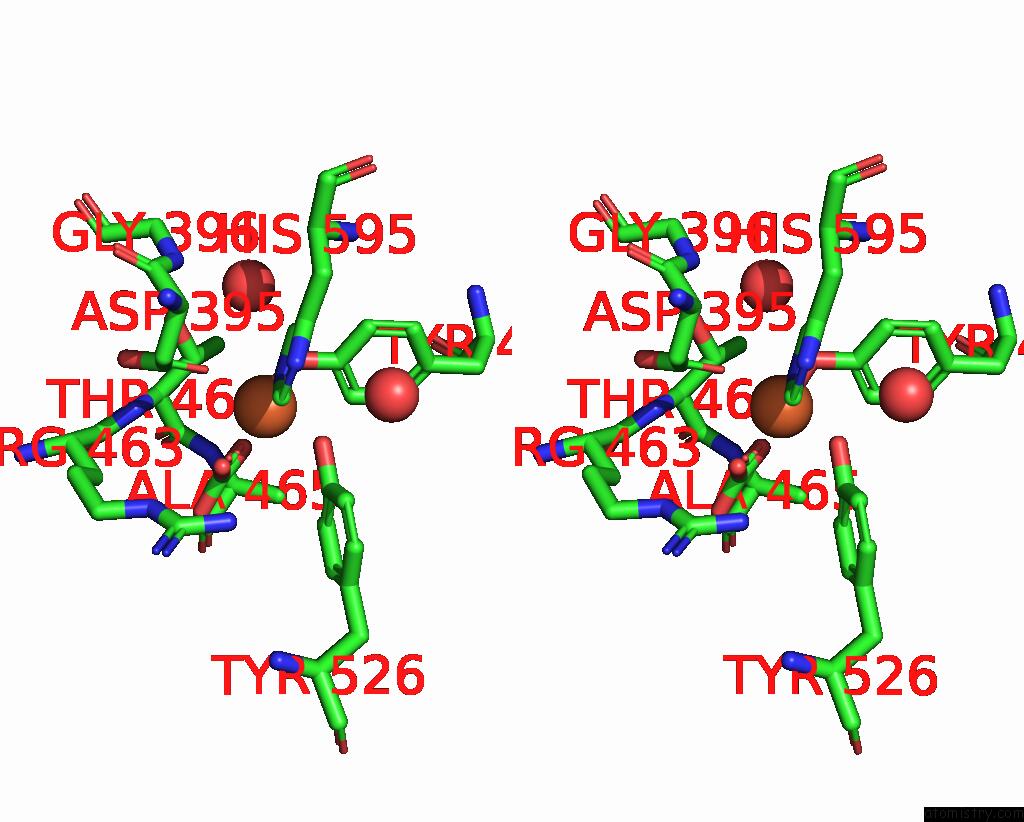
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 1 of Removal of Sugars and Sugars-Like Molecules From the Solution By C- Lobe of Lactoferrin: Crystal Structure of the Complex of C-Lobe with Beta-D-Glucopyranosyl-(1->4)-Beta-D-Galactopyranosyl-(1->4)-Alpha-D- Glucopyranose at 1.9 A Resolution within 5.0Å range:
|
Reference:
R.Mir,
R.P.Kumar,
N.Singh,
G.P.Vikram,
M.Sinha,
A.Bhushan,
P.Kaur,
A.Srinivasan,
S.Sharma,
T.P.Singh.
Specific Interactions of C-Terminal Half (C-Lobe) of Lactoferrin Protein with Edible Sugars: Binding and Structural Studies with Implications on Diabetes. Int.J.Biol.Macromol. V. 47 50 2010.
ISSN: ISSN 0141-8130
PubMed: 20371371
DOI: 10.1016/J.IJBIOMAC.2010.03.021
Page generated: Sun Aug 4 13:41:36 2024
ISSN: ISSN 0141-8130
PubMed: 20371371
DOI: 10.1016/J.IJBIOMAC.2010.03.021
Last articles
Cl in 7ZWYCl in 7ZX4
Cl in 7ZXD
Cl in 7ZX2
Cl in 7ZWX
Cl in 7ZWT
Cl in 7ZWU
Cl in 7ZW2
Cl in 7ZWS
Cl in 7ZWO