Iron »
PDB 5m3l-5mkq »
5mdl »
Iron in PDB 5mdl: Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method
Enzymatic activity of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method
All present enzymatic activity of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method:
1.12.99.6;
1.12.99.6;
Protein crystallography data
The structure of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method, PDB code: 5mdl
was solved by
J.Kalms,
A.Schmidt,
P.Scheerer,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 47.78 / 1.41 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 73.134, 95.568, 119.686, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 13.2 / 16.3 |
Other elements in 5mdl:
The structure of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method also contains other interesting chemical elements:
| Nickel | (Ni) | 1 atom |
| Magnesium | (Mg) | 1 atom |
| Chlorine | (Cl) | 1 atom |
Iron Binding Sites:
Pages:
>>> Page 1 <<< Page 2, Binding sites: 11 - 12;Binding sites:
The binding sites of Iron atom in the Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method (pdb code 5mdl). This binding sites where shown within 5.0 Angstroms radius around Iron atom.In total 12 binding sites of Iron where determined in the Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method, PDB code: 5mdl:
Jump to Iron binding site number: 1; 2; 3; 4; 5; 6; 7; 8; 9; 10;
Iron binding site 1 out of 12 in 5mdl
Go back to
Iron binding site 1 out
of 12 in the Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method
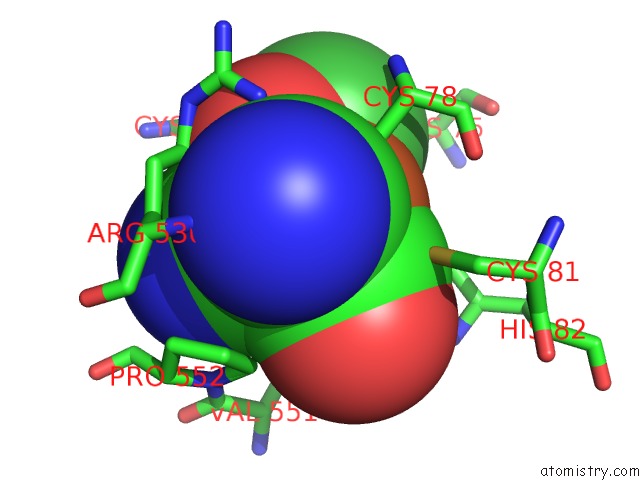
Mono view
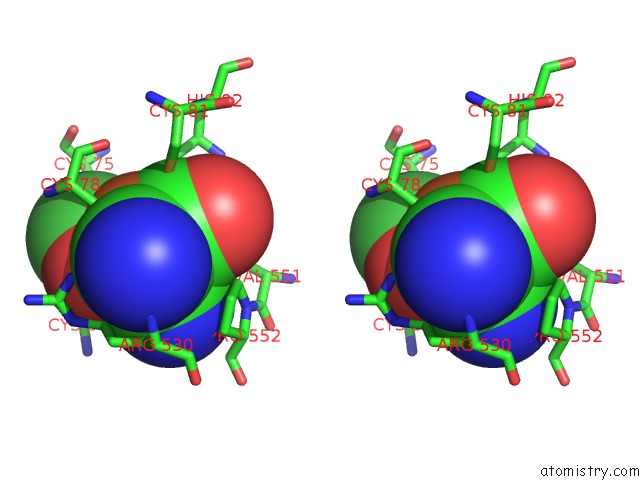
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 1 of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method within 5.0Å range:
|
Iron binding site 2 out of 12 in 5mdl
Go back to
Iron binding site 2 out
of 12 in the Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method
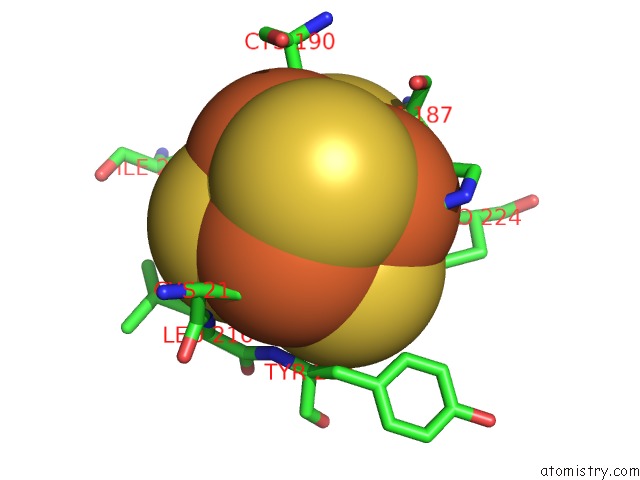
Mono view
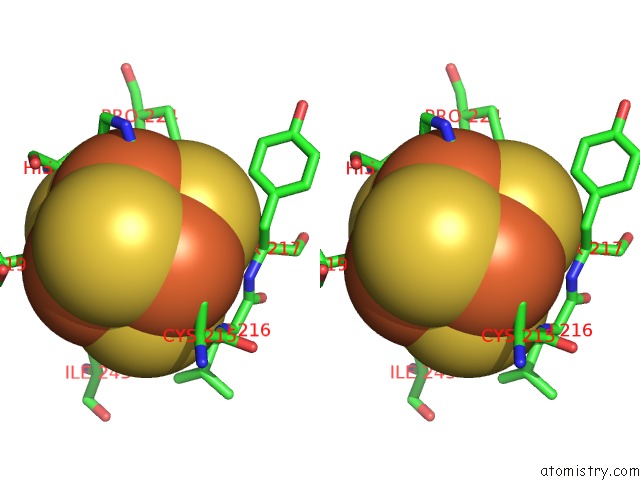
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 2 of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method within 5.0Å range:
|
Iron binding site 3 out of 12 in 5mdl
Go back to
Iron binding site 3 out
of 12 in the Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method
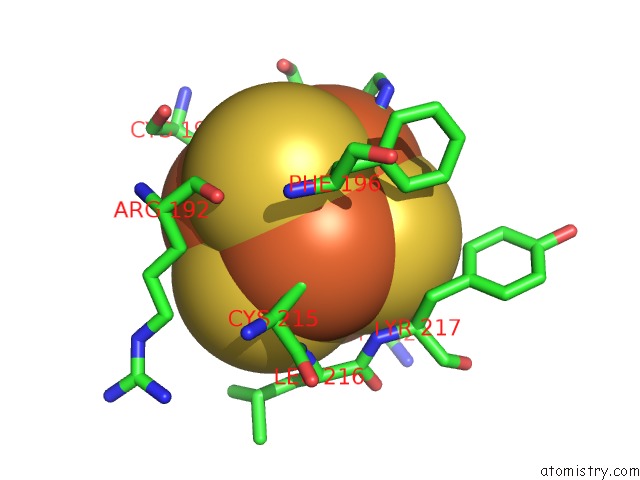
Mono view
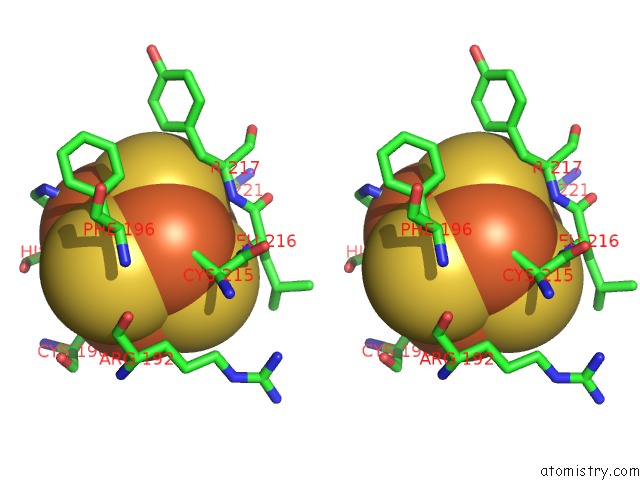
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 3 of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method within 5.0Å range:
|
Iron binding site 4 out of 12 in 5mdl
Go back to
Iron binding site 4 out
of 12 in the Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method
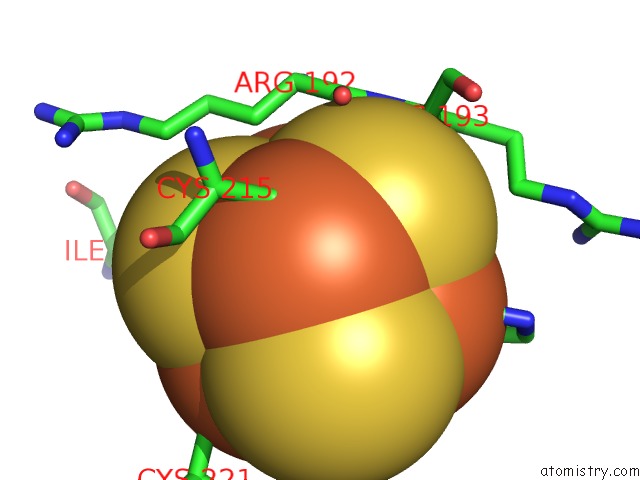
Mono view
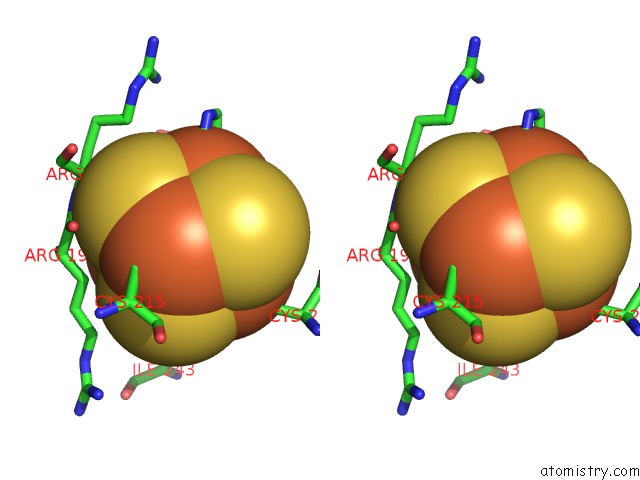
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 4 of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method within 5.0Å range:
|
Iron binding site 5 out of 12 in 5mdl
Go back to
Iron binding site 5 out
of 12 in the Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method
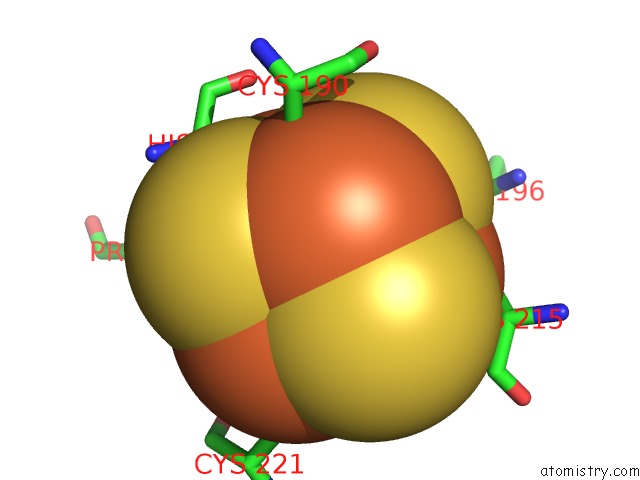
Mono view
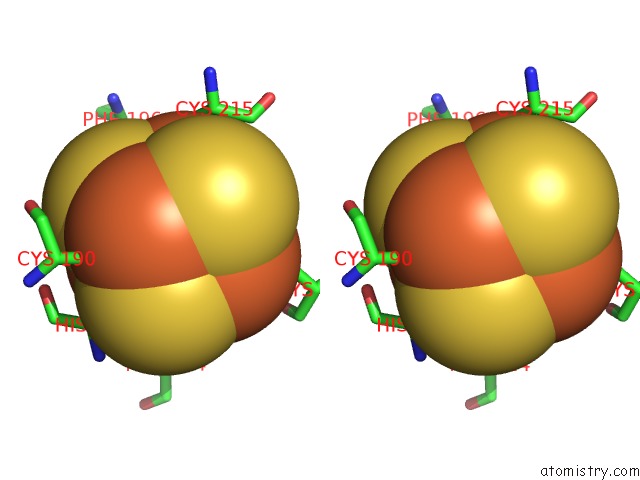
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 5 of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method within 5.0Å range:
|
Iron binding site 6 out of 12 in 5mdl
Go back to
Iron binding site 6 out
of 12 in the Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method
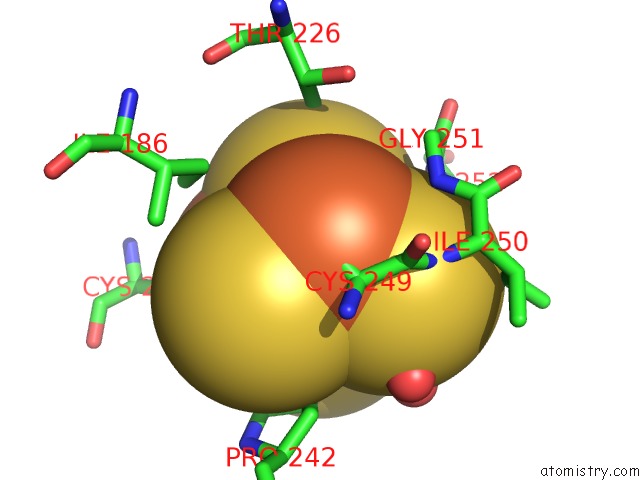
Mono view
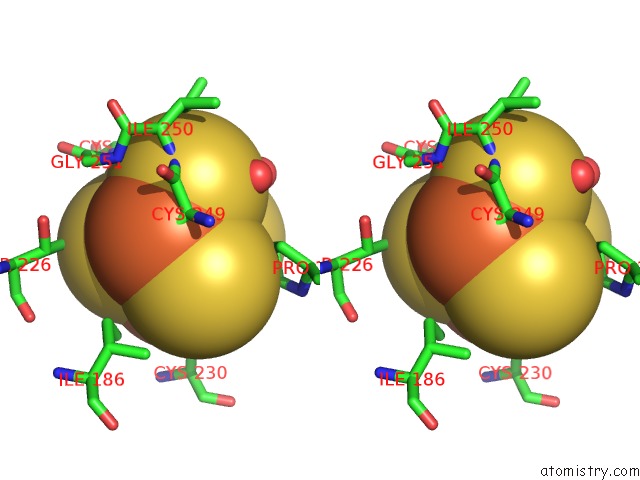
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 6 of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method within 5.0Å range:
|
Iron binding site 7 out of 12 in 5mdl
Go back to
Iron binding site 7 out
of 12 in the Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method
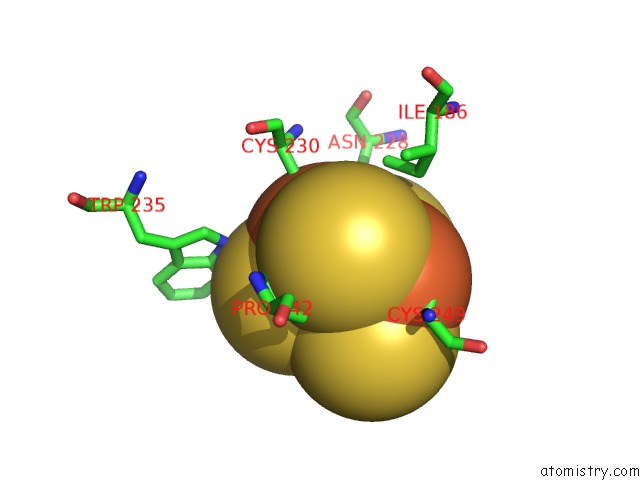
Mono view
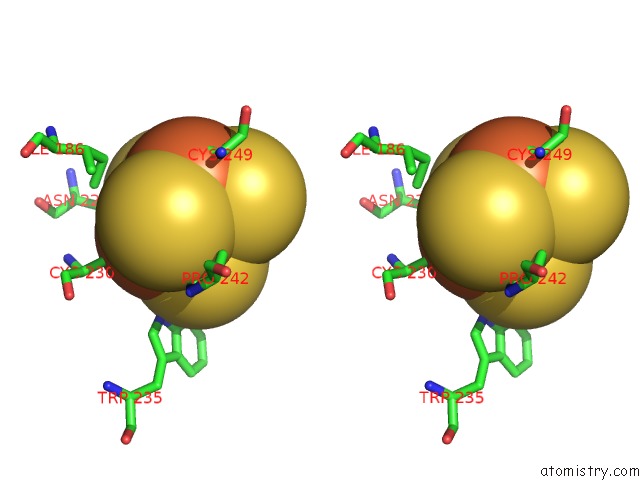
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 7 of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method within 5.0Å range:
|
Iron binding site 8 out of 12 in 5mdl
Go back to
Iron binding site 8 out
of 12 in the Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method
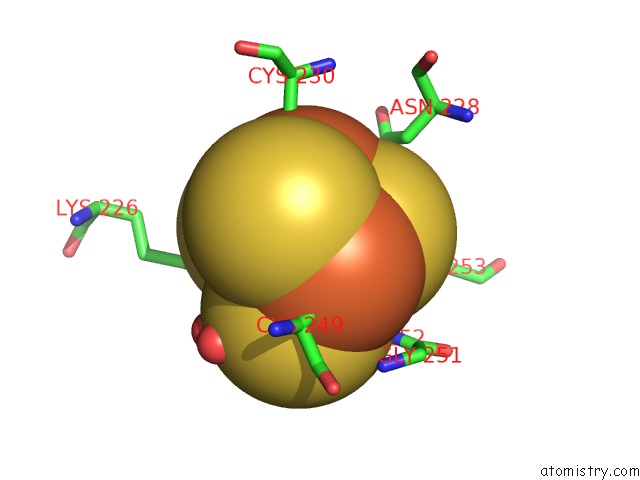
Mono view
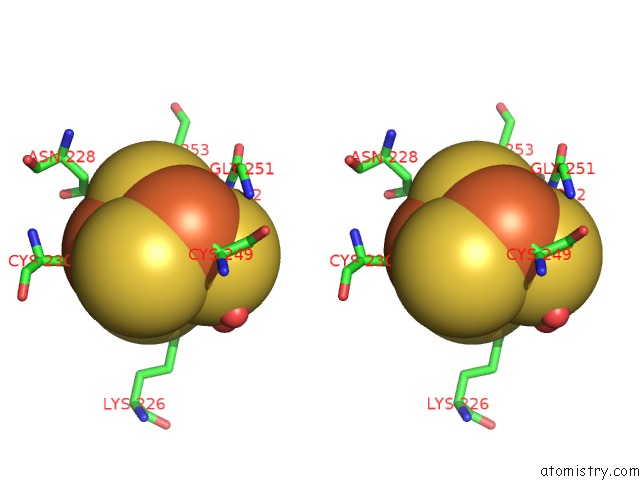
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 8 of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method within 5.0Å range:
|
Iron binding site 9 out of 12 in 5mdl
Go back to
Iron binding site 9 out
of 12 in the Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method
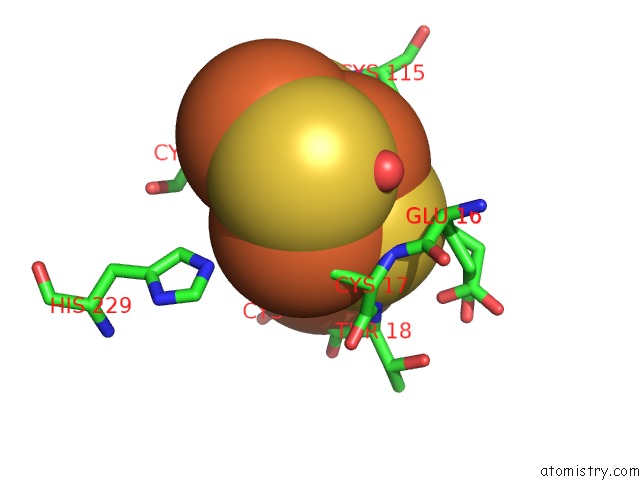
Mono view
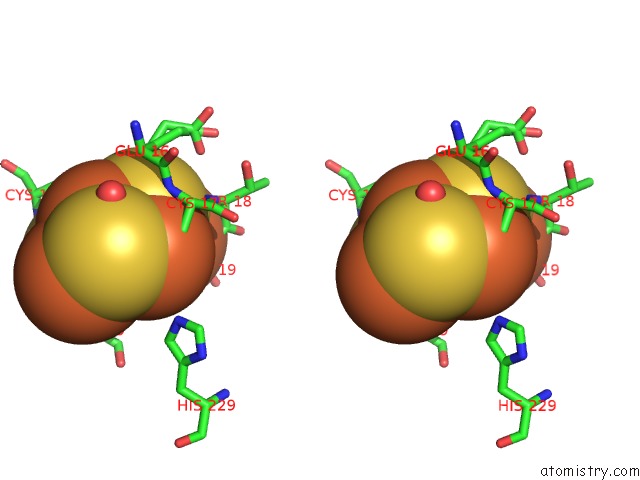
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 9 of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method within 5.0Å range:
|
Iron binding site 10 out of 12 in 5mdl
Go back to
Iron binding site 10 out
of 12 in the Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method
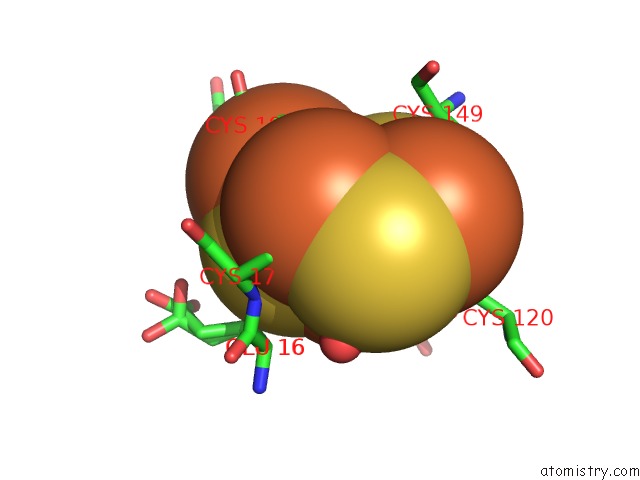
Mono view
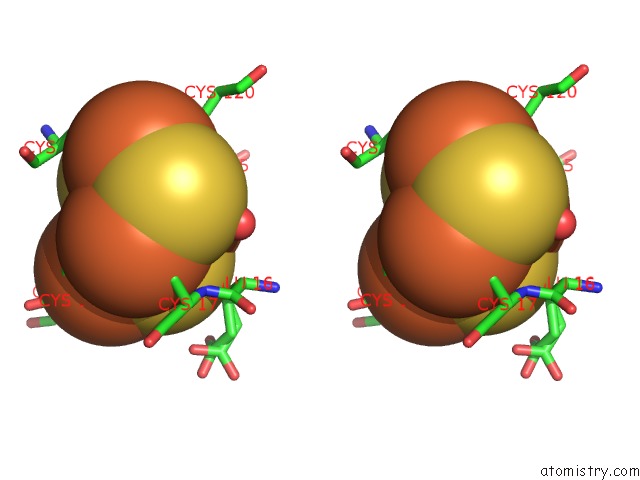
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iron with other atoms in the Fe binding
site number 10 of Crystal Structure of An O2-Tolerant [Nife]-Hydrogenase From Ralstonia Eutropha in Its O2-Derivatized Form By A "Soak-and-Freeze" Derivatization Method within 5.0Å range:
|
Reference:
J.Kalms,
A.Schmidt,
S.Frielingsdorf,
T.Utesch,
G.Gotthard,
D.Von Stetten,
P.Van Der Linden,
A.Royant,
M.A.Mroginski,
P.Carpentier,
O.Lenz,
P.Scheerer.
Tracking the Route of Molecular Oxygen in O2-Tolerant Membrane-Bound [Nife] Hydrogenase. Proc. Natl. Acad. Sci. V. 115 E2229 2018U.S.A..
ISSN: ESSN 1091-6490
PubMed: 29463722
DOI: 10.1073/PNAS.1712267115
Page generated: Tue Aug 5 23:53:12 2025
ISSN: ESSN 1091-6490
PubMed: 29463722
DOI: 10.1073/PNAS.1712267115
Last articles
Fe in 6WZ1Fe in 6WZ0
Fe in 6WYH
Fe in 6WYG
Fe in 6WYU
Fe in 6WYD
Fe in 6WYF
Fe in 6WY7
Fe in 6WY0
Fe in 6WXU